***
Note: Updated CRACR information and scripts can be found here
***
Inferring condition-specific
transcription factor function
from DNA binding and gene expression
data
This website supports
McCord, et al.
Molecular Systems Biology, 3:100
Numerous genomic and proteomic datasets are
permitting the elucidation of transcriptional regulatory networks in the yeast
Saccharomyces cerevisiae. However, predicting the condition dependence of
regulatory network interactions has been challenging, because most
protein–DNA interactions identified in vivo are from assays performed in
one or a few cellular states. Here, we present a novel method to predict the
condition-specific functions of S. cerevisiae transcription factors (TFs) by
integrating 1327 microarray gene expression data sets and either comprehensive
TF binding site data from protein
binding microarrays (PBMs) or in silico motif data. Importantly, our
method does not impose arbitrary thresholds for calling target regions
‘bound’ or genes ‘differentially expressed’, but rather
allows all the information derived from a TF binding or gene expression
experiment to be considered. We show that this method can identify
environmental, physical, and genetic interactions, as well as distinct sets of
genes that might be activated or repressed by a single TF under particular
conditions. This approach can be used to suggest conditions for directed in
vivo experimentation and to predict TF function.
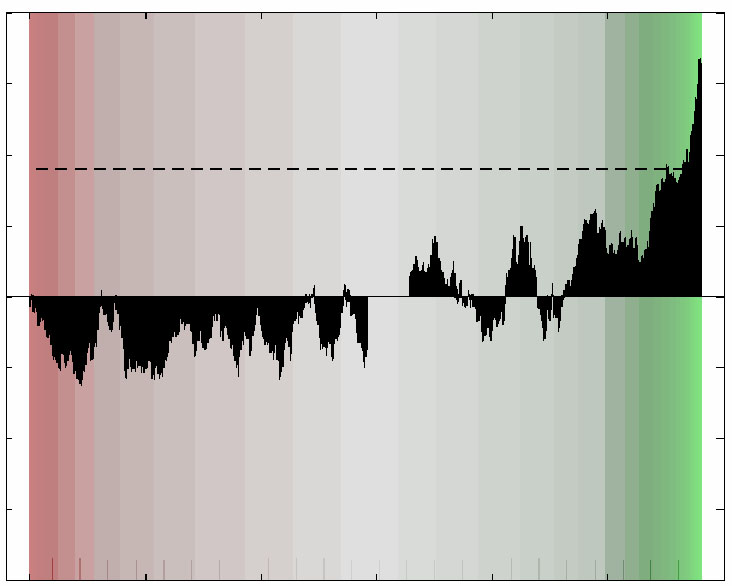
Below are Supplementary data files that accompany this manuscript which do not appear in the Supplementary Information on the MSB website.
CRACR algorithm: *coming soon* This zip file contains all the components necessary to run CRACR. Requires MATLAB and Perl. This zip file also includes includes properly formatted sample datafiles (PBM and gene expression data) that can be used in conjunction with the CRACR algorithm to generate some of the figures included in the manuscript.
CRACR readme: *coming soon* A text file explaining how to use the files above to run the CRACR algorithm.
Expression
Data: A tab-delimited file containing data from all 1327 expression
conditions used in this publication.
All data is presented as log2 expression ratios. Genes with no data in a particular
expression experiment are assigned an expression value of “
Expression Condition Annotations: An Excel file containing the condition annotation terms for all 1327 expression conditions.
Questions? Rachel Patton McCord, Mike Berger, Anthony Philippakis, Martha Bulyk.